The detection pipeline already returned result === 'not_vine' but the app rendered it the same as a low-confidence positive, which was confusing (a coffee cup classified at 35% would show up as "uncertain vine"). Surface non-vine results explicitly across the app: - New ScanStatus 'not_vine' branch in types/detection.getScanStatus() - StatusTag, ScanListItem fill, MapBottomSheet row icon (HelpCircle) and MapView marker color get a neutral grey palette for not_vine - ResultScreen short-circuits to a centered "Aucune vigne détectée" layout with a single CTA "Reprendre une photo" (instead of pretending the model has a meaningful prediction to show) - MapBottomSheet learns an isLoading prop and renders 4 row skeletons while useHistory rehydrates, instead of flashing the "no plants" empty state. MapScreen plumbs historyLoading through Bundles the i18n additions (FR + EN) for this commit and the next two: result.notVineTitle/Message, myPlants.status.notVine, plus the network.* and scanner.galleryComingSoon* keys used by follow-up commits — splitting JSON hunks would have been more churn than signal. Co-Authored-By: Claude Opus 4.7 (1M context) <noreply@anthropic.com> |
||
|---|---|---|
| .claude | ||
| docs | ||
| venv | ||
| VinEye | ||
| vineye-admin | ||
| .gitignore | ||
| AGENTS.md | ||
| CLAUDE.md | ||
| package.json | ||
| pnpm-lock.yaml | ||
| PROJECT_SUMMARY.md | ||
| README.md | ||
Tensorflow Grapevine Disease Detection
Description
This project develops a mobile application for detecting diseases on grapevines using a Deep Learning model. The implementation leverages TensorFlow and Keras to build a CNN-based classifier for identifying three common diseases: Black Rot, ESA (Net Blight), and Leaf Blight.
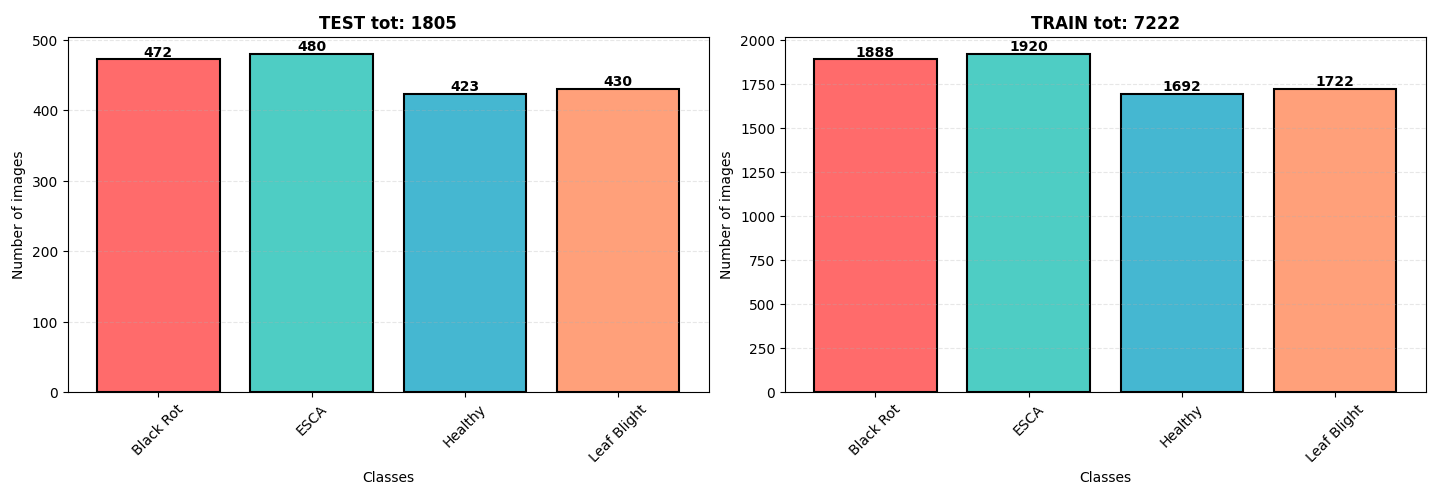
📁 Dataset
The dataset originates from Kaggle, containing 9,027 images of grapevine leaves. The diseases are categorized as:
- Black Rot
- ESCA (Net Blight)
- Leaf Blight
The dataset is well-balanced with a slight overrepresentation of ESCA and Black Rot. All images are in .jpeg format with dimensions 256x256 pixels.
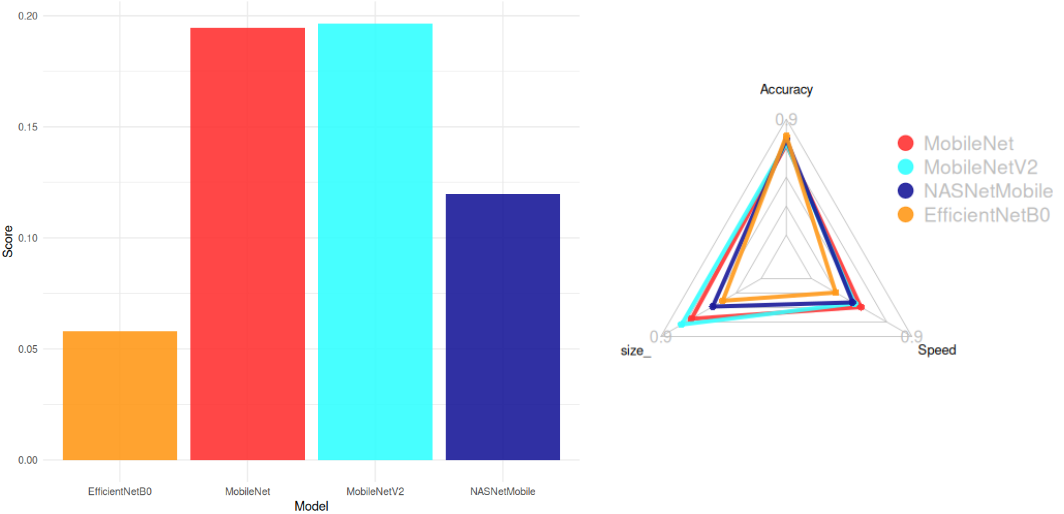
📊 Model Architecture Selection
We evaluated pre-trained models from keras applications to balance accuracy, model size, and inference speed. The selection criteria included:
- Maximize accuracy
- Minimize size (1/size)
- Maximize CPU speed (1/CPU Time)
The score formula used for selection was:
Score = \frac{Accuracy}{Size . CPU Time}
Top Models (Score > 0.05):
- MobileNetV2 (Smallest: 14 MB, High Accuracy: 77%)
- MobileNet (Fastest: 22.6 ms)
- NASNetMobile
- EfficientNetB0
Conclusion:
MobileNetV2 was chosen for its optimal balance between accuracy, size, and speed.
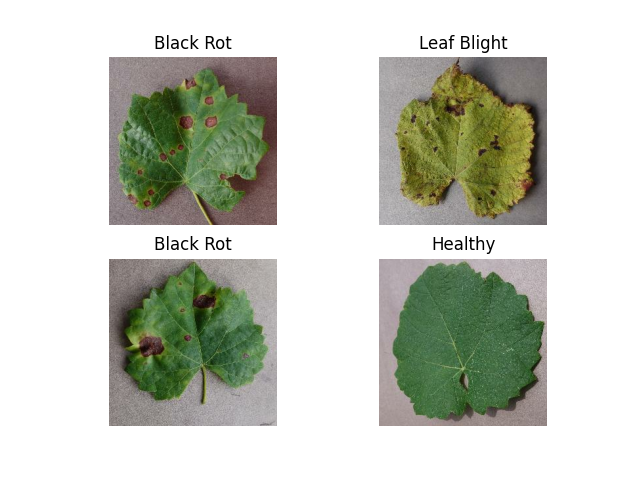
🍇 Grapevine Diseases
Key Diseases:
- Black Rot
- ESCA (Net Blight)
- Leaf Blight
🤖 Model Structure
Architecture:
# Auto Stop
early_stopping = EarlyStopping(monitor="val_loss", min_delta=0.2, patience=10)
# Model
model = Sequential()
model.add(tf.keras.applications.MobileNetV2(
input_shape=(IMG_HEIGHT, IMG_WIDTH, CHANNELS),
include_top=False,
weights='imagenet'
))
model.add(tf.keras.layers.GlobalAveragePooling2D())
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(NUM_CLASSES, activation='softmax'))
optimizer = tf.keras.optimizers.Adam(learning_rate=LEARNING_RATE)
model.compile(
optimizer=optimizer,
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy']
)
Parameters:
- Total params: 7.12M (27.17 MB)
- Trainable params: 2.36M (9.01 MB)
- Non-trainable params: 34.11K (133.25 KB)
🛠️ Training Details
- Batch Size: 32
- Epochs: 100 (reduced to 25 via early stopping)
- Data Augmentation: Not used (insufficient improvement in accuracy)
- Normalization: Pixel values normalized to [0, 1]
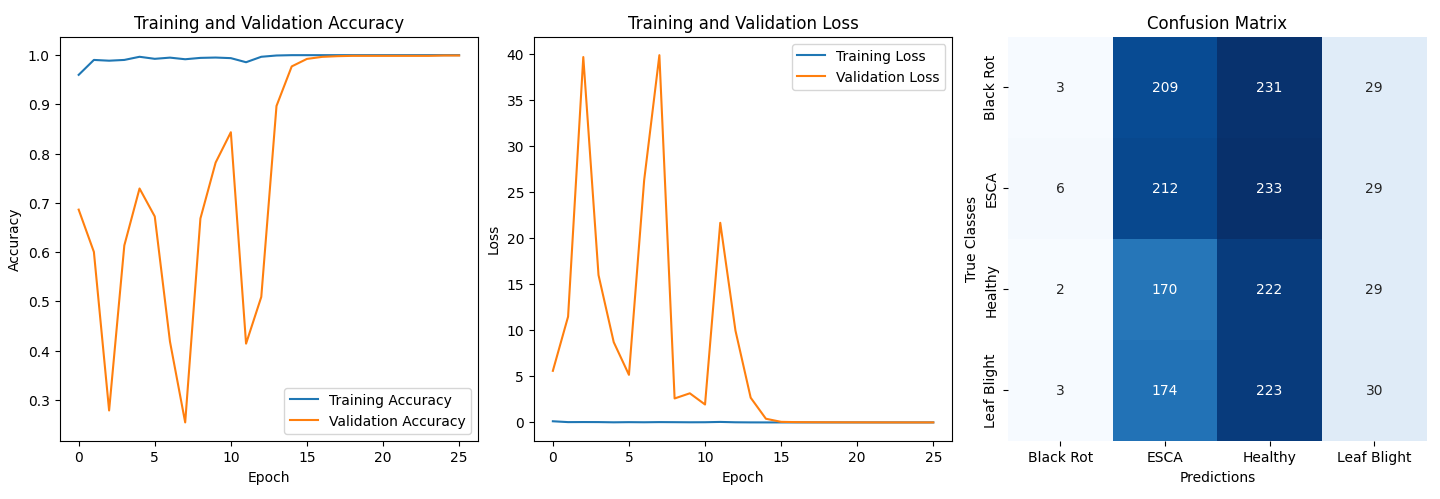
📊 Results
Performance:
- Validation Accuracy: ~99.9%
- Confusion Matrix Analysis:
- Model biased toward ESCA and Healthy classes.
- Suspected causes:
- Original dataset imbalance
- Similar visual features across diseases
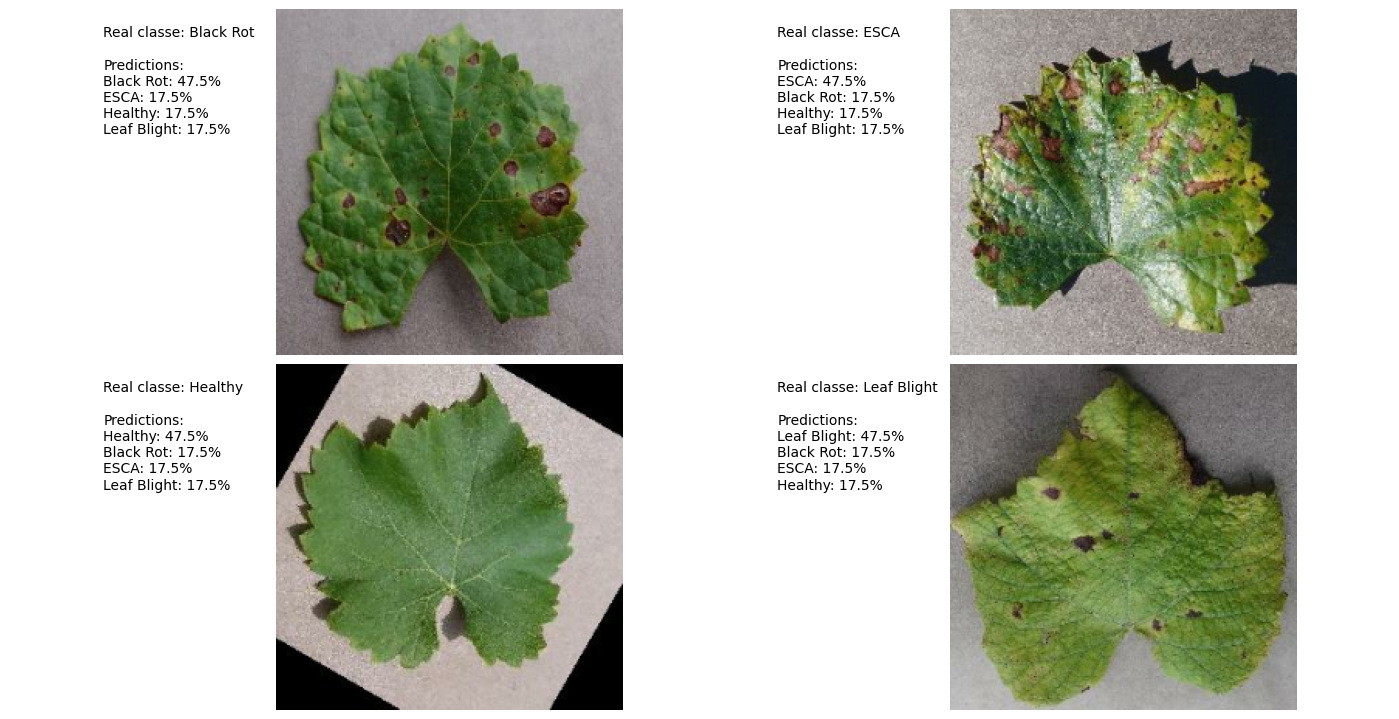
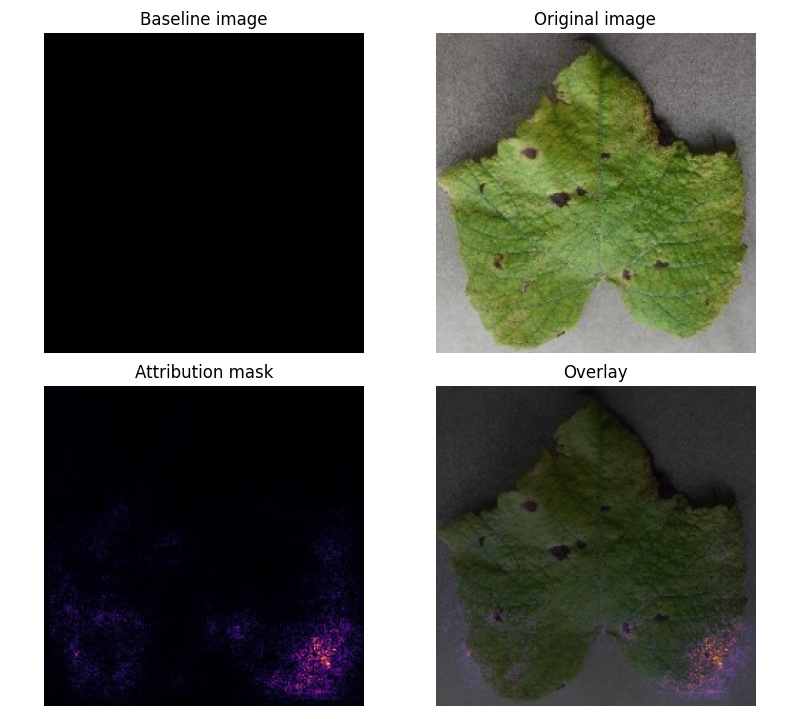
Prediction Example:
Attribution Mask:
- Key Insight: Model focuses on leaf shape rather than disease-specific features (e.g., black spots).
📚 ressources:
https://www.kaggle.com/code/ahmedmsaber/grape-leafs-diseases-mobilenetv2-val-acc-99
https://www.tensorflow.org/tutorials/images/classification?hl=en
https://www.tensorflow.org/lite/convert?hl=en
https://www.tensorflow.org/tutorials/interpretability/integrated_gradients?hl=en
🤖AI(s) : deepseek-coder:6.7b | deepseek-r1:8b