New screens: Guides, Library, Map, Notifications, Settings. Home refactored with modular components (SearchHeader, HomeCta, FrequentDiseases, SeasonAlert, PracticalGuides). Replaced hardcoded colors with theme tokens in SeasonAlert and HomeCta. Updated navigation, i18n, and CLAUDE.md. Co-Authored-By: Claude Opus 4.6 (1M context) <noreply@anthropic.com> |
||
|---|---|---|
| docs/images | ||
| venv | ||
| VinEye | ||
| .gitignore | ||
| AGENTS.md | ||
| README.md | ||
Tensorflow Grapevine Disease Detection
This document outlines the development of a modile application that uses a DeepLearning model de detect diseases on grapevine.
Dataset
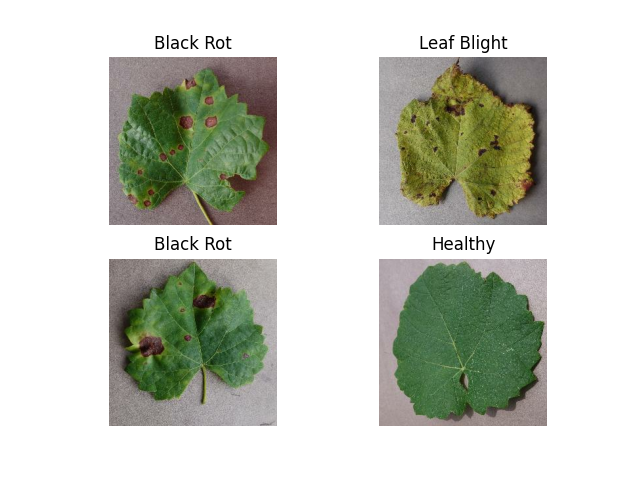
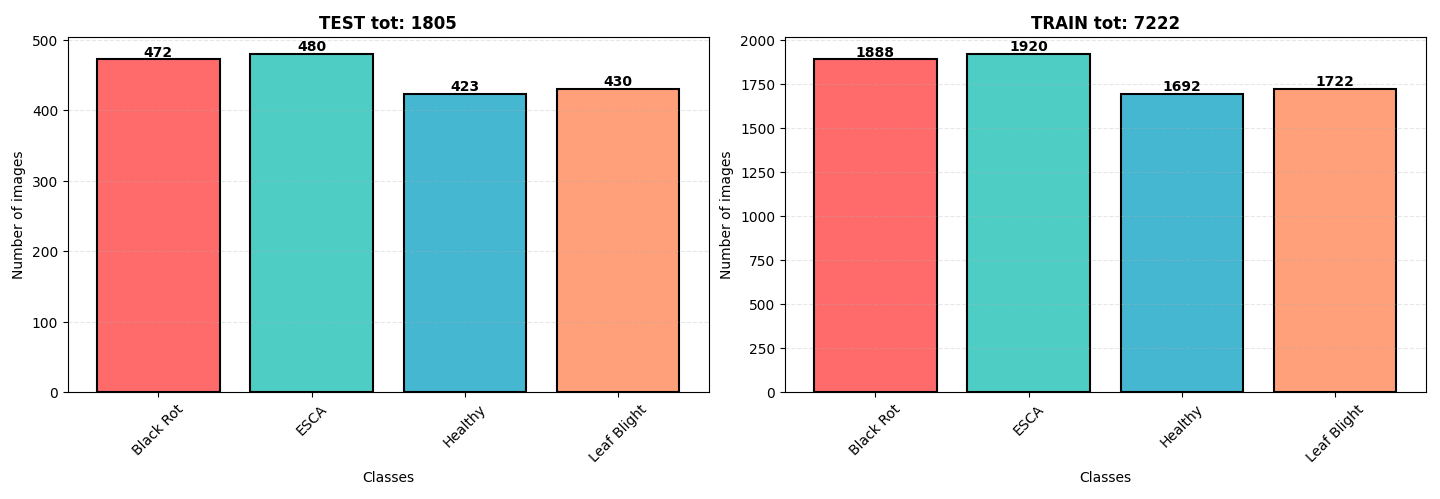
The data used in this study came from kaggle. It is split into training, validation, and testing sets ensuring a robust evaluation of our model's performance. The dataset consists of a set of 9027 images of three disease commonly found on grapevines: Black Rot, ESCA, and Leaf Blight. Classes are well balenced with a slit overrepresentation of ESCA and Black Rot . Images are in .jpeg format with dimensions of 256x256 pixels.
Model Structure
Our model is a Convolutional Neural Network (CNN) built using Keras API with TensorFlow backend. It includes several convolutional layers followed by batch normalization, ReLU activation function and max pooling for downsampling. Dropout layers are used for regularization to prevent overfitting. The architecture details and parameters are as follows:
model = Sequential([
data_augmentation,
# Block 1
layers.Conv2D(32, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.Conv2D(32, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.MaxPooling2D(pool_size=2),
layers.Dropout(0.25),
# Block 2
layers.Conv2D(64, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.Conv2D(64, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.MaxPooling2D(pool_size=2),
layers.Dropout(0.25),
# Block 3
layers.Conv2D(128, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.Conv2D(128, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.MaxPooling2D(pool_size=2),
layers.Dropout(0.25),
# Block 4
layers.Conv2D(256, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.Conv2D(256, kernel_size=3, padding='same', activation='relu'),
layers.BatchNormalization(),
layers.MaxPooling2D(pool_size=2),
layers.Dropout(0.25),
# Classification head
layers.GlobalAveragePooling2D(),
layers.Dense(256, activation='relu'),
layers.BatchNormalization(),
layers.Dropout(0.5),
layers.Dense(128, activation='relu'),
layers.BatchNormalization(),
layers.Dropout(0.5),
layers.Dense(num_classes)
])
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
Total params: 3,825,134 (14.59 MB)
Trainable params: 1,274,148 (4.86 MB)
Non-trainable params: 2,688 (10.50 KB)
Optimizer params: 2,548,298 (9.72 MB)
Training Details
Training was done using a batch size of 32 over 100 epochs. Data augmentation methods include horizontal/vertical flip (RandomFlip), rotation (RandomRotation), zooming (RandomZoom) and rescaling (Rescaling). Pixel values are normalized to the range [0, 1] after loading.
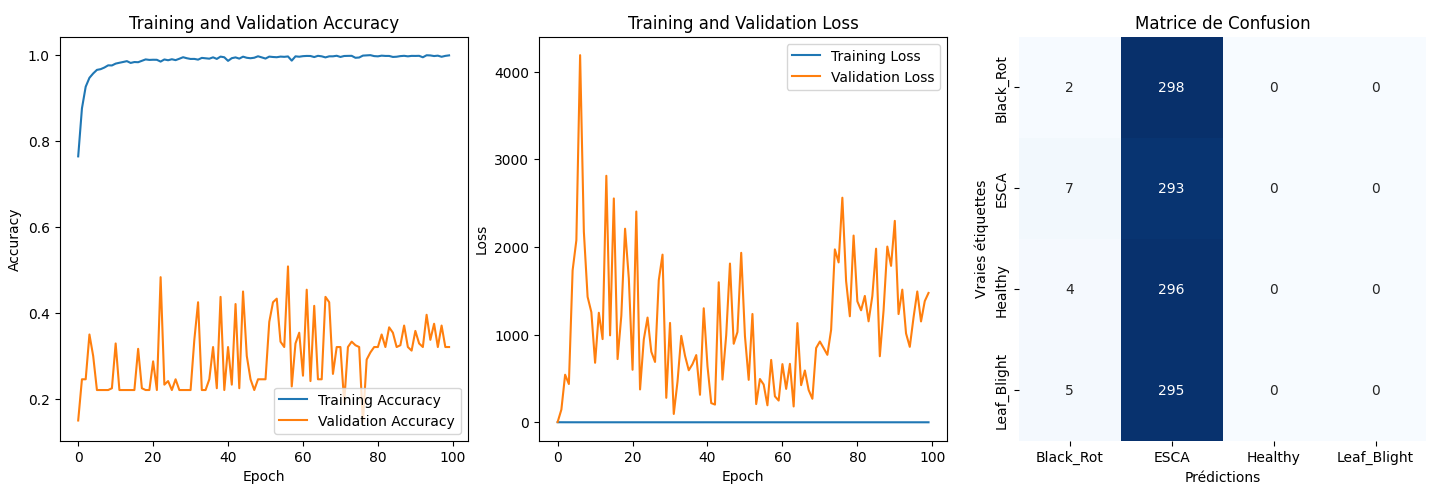
Results
Our best model's performance has an average accuracy of roughly 30% on the validation set. This suggests potential overfitting towards the ESCA class. However, the model can identify key features that distinguish all classes: marks on the leaves (fig.4).
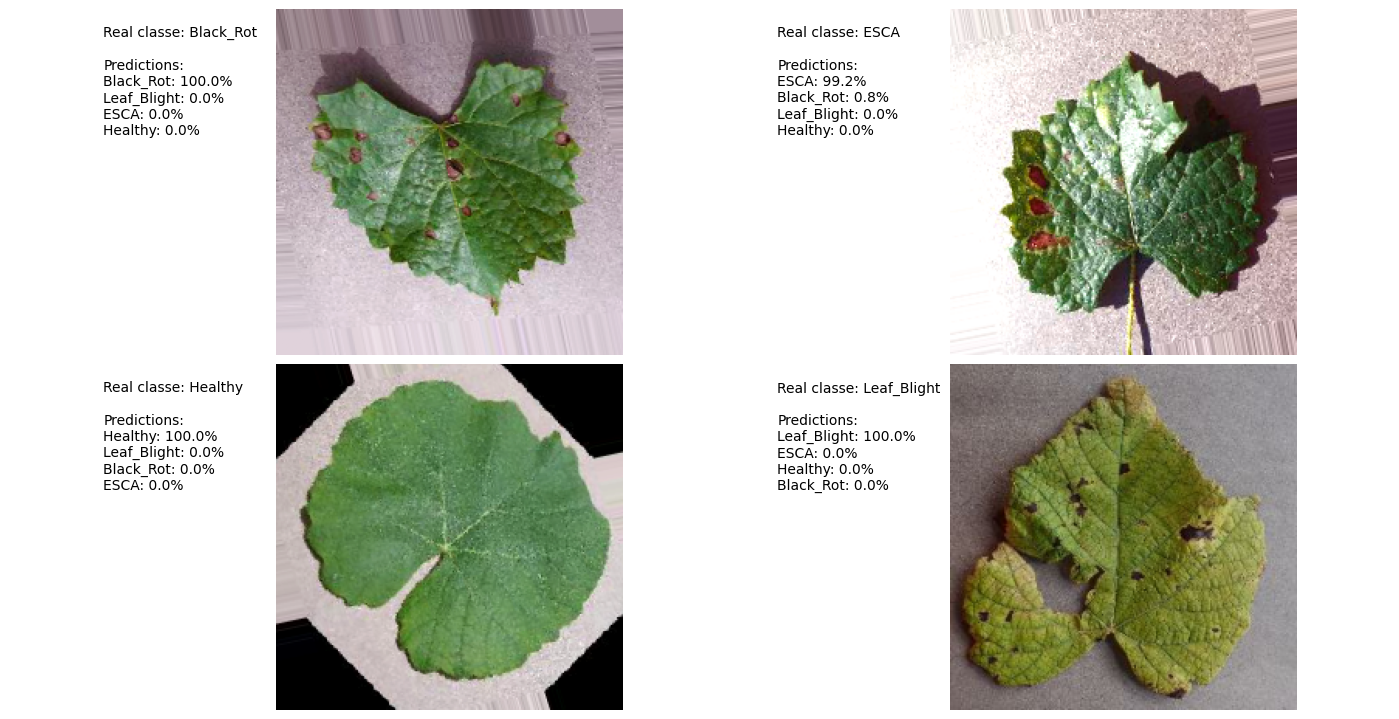
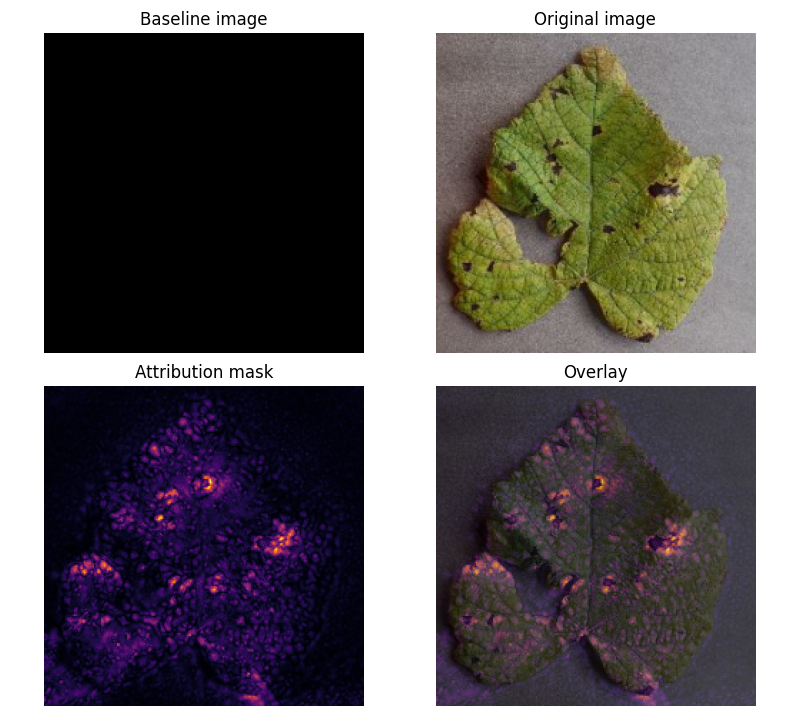
Prediction Example
Attribution Mask
The attribution mask provides an insight into what features the model has learned to extract from each image, which can be seen in figure 4. This can help guide future work on improving disease detection and understanding how the model is identifying key features for accurate classification.
ressources:
https://www.tensorflow.org/tutorials/images/classification?hl=en
https://www.tensorflow.org/lite/convert?hl=en
https://www.tensorflow.org/tutorials/interpretability/integrated_gradients?hl=en
AI(s) : deepseek-coder:6.7b | deepseek-r1:8b