Touch targets on the bottom tabs were too small near the top edge of the bar, making it easy to miss the tap when reaching for an icon. Add hitSlop on the Pressable and pointerEvents="none" on the active-state badge so taps always land on the parent button. Co-Authored-By: Claude Opus 4.7 (1M context) <noreply@anthropic.com> |
||
|---|---|---|
| .claude | ||
| docs | ||
| venv | ||
| VinEye | ||
| vineye-admin | ||
| .gitignore | ||
| AGENTS.md | ||
| CLAUDE.md | ||
| package.json | ||
| pnpm-lock.yaml | ||
| PROJECT_SUMMARY.md | ||
| README.md | ||
Tensorflow Grapevine Disease Detection
Description
This project develops a mobile application for detecting diseases on grapevines using a Deep Learning model. The implementation leverages TensorFlow and Keras to build a CNN-based classifier for identifying three common diseases: Black Rot, ESA (Net Blight), and Leaf Blight.
📁 Dataset
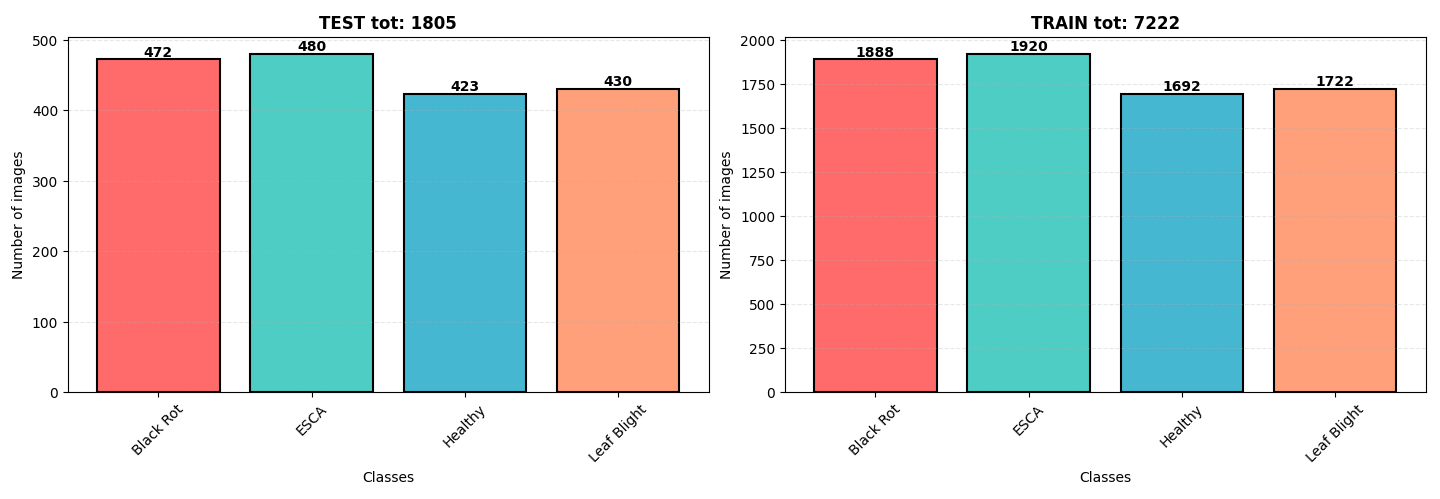
The dataset originates from Kaggle, containing 9,027 images of grapevine leaves. The diseases are categorized as:
- Black Rot
- ESCA (Net Blight)
- Leaf Blight
The dataset is well-balanced with a slight overrepresentation of ESCA and Black Rot. All images are in .jpeg format with dimensions 256x256 pixels.
📊 Model Architecture Selection
We evaluated pre-trained models from keras applications to balance accuracy, model size, and inference speed. The selection criteria included:
- Maximize accuracy
- Minimize size (1/size)
- Maximize CPU speed (1/CPU Time)
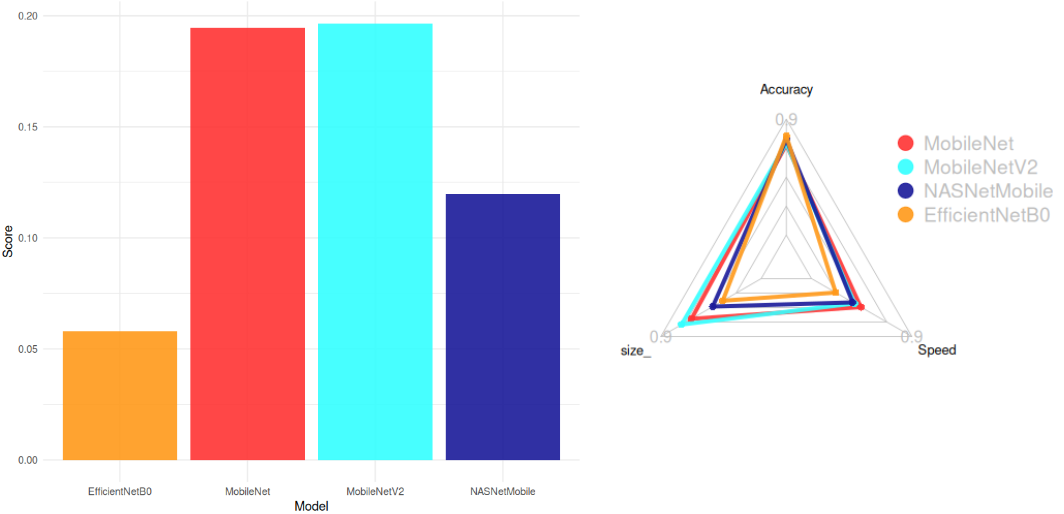
The score formula used for selection was:
Score = \frac{Accuracy}{Size . CPU Time}
Top Models (Score > 0.05):
- MobileNetV2 (Smallest: 14 MB, High Accuracy: 77%)
- MobileNet (Fastest: 22.6 ms)
- NASNetMobile
- EfficientNetB0
Conclusion:
MobileNetV2 was chosen for its optimal balance between accuracy, size, and speed.
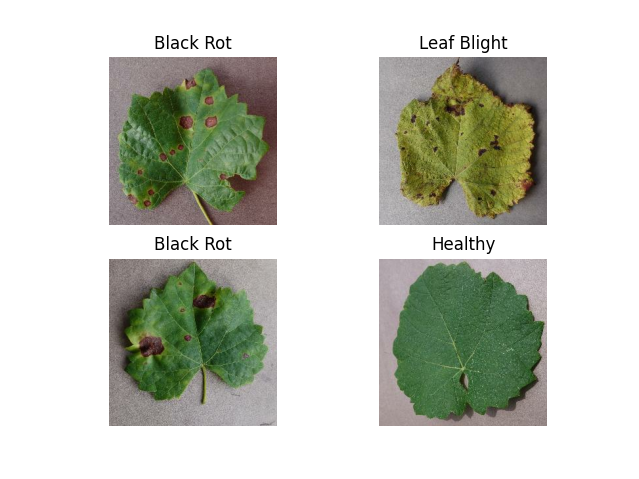
🍇 Grapevine Diseases
Key Diseases:
- Black Rot
- ESCA (Net Blight)
- Leaf Blight
🤖 Model Structure
Architecture:
# Auto Stop
early_stopping = EarlyStopping(monitor="val_loss", min_delta=0.2, patience=10)
# Model
model = Sequential()
model.add(tf.keras.applications.MobileNetV2(
input_shape=(IMG_HEIGHT, IMG_WIDTH, CHANNELS),
include_top=False,
weights='imagenet'
))
model.add(tf.keras.layers.GlobalAveragePooling2D())
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(NUM_CLASSES, activation='softmax'))
optimizer = tf.keras.optimizers.Adam(learning_rate=LEARNING_RATE)
model.compile(
optimizer=optimizer,
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy']
)
Parameters:
- Total params: 7.12M (27.17 MB)
- Trainable params: 2.36M (9.01 MB)
- Non-trainable params: 34.11K (133.25 KB)
🛠️ Training Details
- Batch Size: 32
- Epochs: 100 (reduced to 25 via early stopping)
- Data Augmentation: Not used (insufficient improvement in accuracy)
- Normalization: Pixel values normalized to [0, 1]
📊 Results
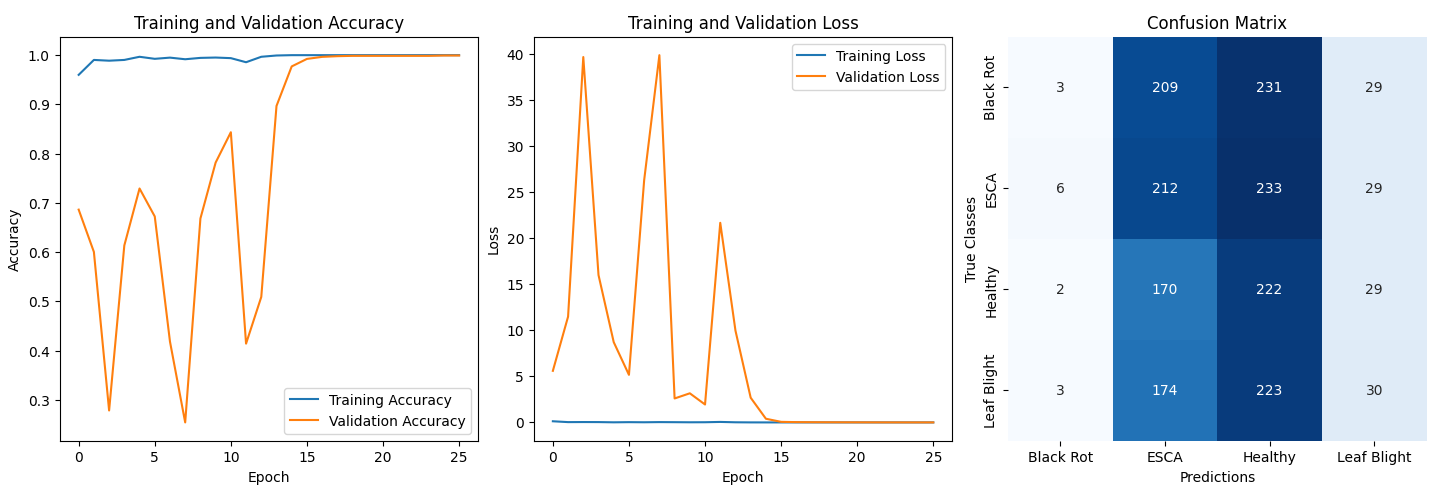
Performance:
- Validation Accuracy: ~99.9%
- Confusion Matrix Analysis:
- Model biased toward ESCA and Healthy classes.
- Suspected causes:
- Original dataset imbalance
- Similar visual features across diseases
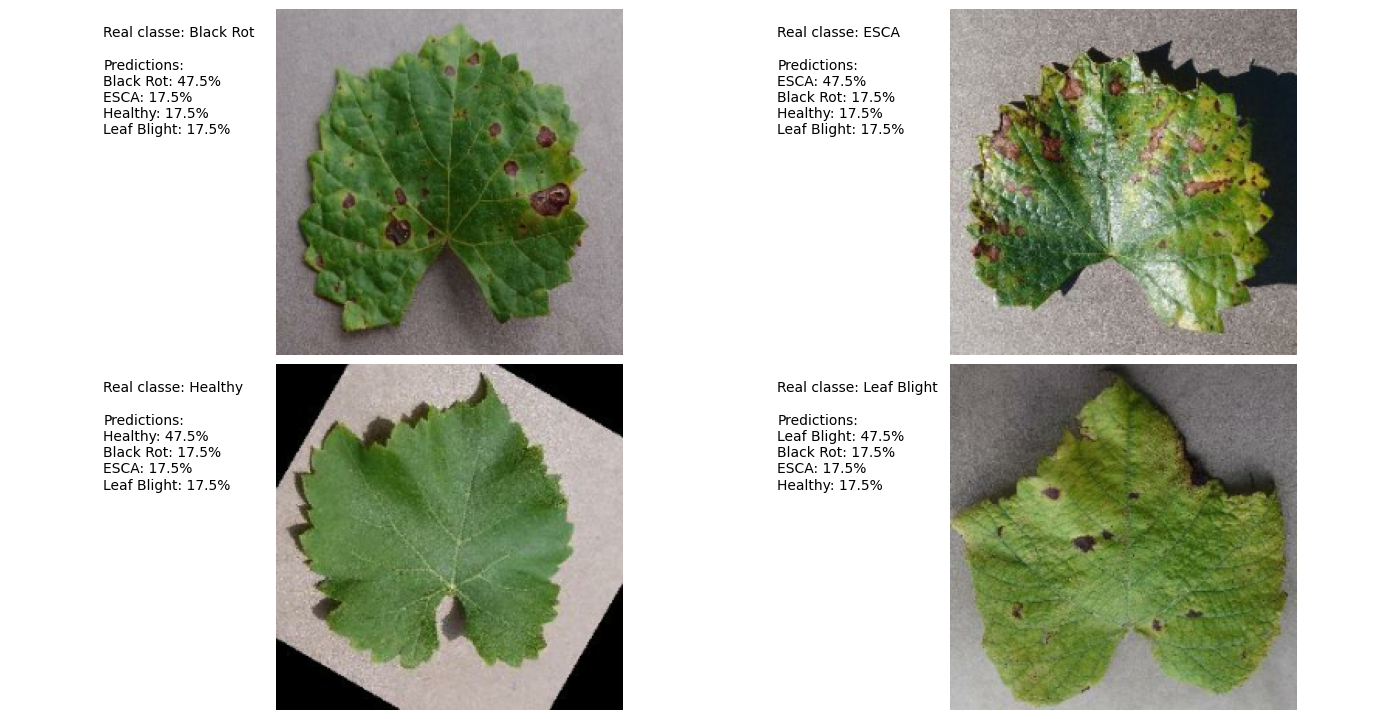
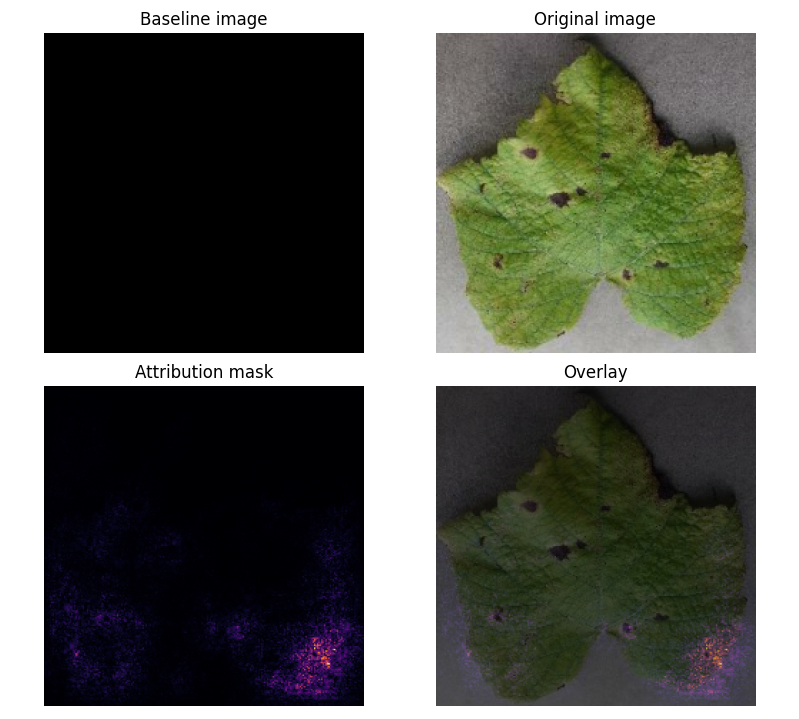
Prediction Example:
Attribution Mask:
- Key Insight: Model focuses on leaf shape rather than disease-specific features (e.g., black spots).
📚 ressources:
https://www.kaggle.com/code/ahmedmsaber/grape-leafs-diseases-mobilenetv2-val-acc-99
https://www.tensorflow.org/tutorials/images/classification?hl=en
https://www.tensorflow.org/lite/convert?hl=en
https://www.tensorflow.org/tutorials/interpretability/integrated_gradients?hl=en
🤖AI(s) : deepseek-coder:6.7b | deepseek-r1:8b