Une seule SearchScreen modale gère la recherche depuis Home, MyPlants,
Guides et Map. La SearchBar partagée passe en mode trigger sur ces pages,
ouvrant la modal au tap.
Animation par contexte (native-stack) :
- Home/MyPlants/Guides → 'fade_from_bottom' (fade + léger glissement)
- Map → 'fade' pur (la barre est déjà en haut, pas de mouvement vertical)
via param de route { fromMap: true }
SearchScreen — mode global :
- 3 catégories : Maladies (red), Guides pratiques (blue), Mes plantes (green)
- Filter chips scrollable horizontal : Tout / Maladies / Guides / Plantes
avec count par catégorie
- Sections en accordion (chevron) avec count par section
- Tag coloré sur chaque résultat indiquant la catégorie
- Plantes scannées incluses (recherche par customName / cépage)
- Tap résultat : DiseaseDetail / GuideDetail / Map+focusScanId / ScanDetail
SearchScreen — mode Map (fromMap=true) :
- Liste plate des plantes localisées uniquement
- Distance haversine depuis position courante (formatDistance helper)
- Tri par distance croissante
- Tap → navigate Main/Map avec focusScanId
MapScreen :
- useEffect sur route.params?.focusScanId : animation 2 étapes (zoom large
450ms → zoom serré 500ms) + ouverture du preview
- Caméra décalée vers le sud (lat - delta * 0.18) pour que le marker soit
visible AU-DESSUS du bottom sheet, pas masqué dessous
- Reset du param après usage via setParams
Recents :
- Hook useRecentSearches (AsyncStorage @vineye:recent_searches)
- Max 10, déduplication insensible à la casse
- Affichés quand pas de query, avec clear all + remove individuel
SearchBar partagée :
- Refactorisée 100% Tailwind (sauf 2-3 props RN-spécifiques sur TextInput)
- Nouvelle prop onTriggerPress : devient un Pressable avec Text placeholder
qui navigate vers Search au lieu d'être un input local
Wirings :
- SearchSection (Home + Guides) : trigger
- MyPlantsScreen : trigger (suppression de l'inline filtering, plus utilisé)
- FloatingSearch (Map) : trigger avec fromMap: true
- RootNavigator : Stack.Screen Search avec options dynamique selon param
i18n FR + EN :
- search.{placeholder, placeholderMap, recentTitle, clearAll, noRecent,
resultsTitle, noResults, nearbyPlantsTitle, noPlants}
- search.filter.{all, diseases, guides, plants}
- search.section.{diseases, guides, plants}
- search.tag.{disease, guide, plant}
Co-Authored-By: Claude Opus 4.7 (1M context) <noreply@anthropic.com>
|
||
|---|---|---|
| .claude | ||
| docs | ||
| venv | ||
| VinEye | ||
| vineye-admin | ||
| .gitignore | ||
| AGENTS.md | ||
| CLAUDE.md | ||
| package.json | ||
| pnpm-lock.yaml | ||
| PROJECT_SUMMARY.md | ||
| README.md | ||
Tensorflow Grapevine Disease Detection
Description
This project develops a mobile application for detecting diseases on grapevines using a Deep Learning model. The implementation leverages TensorFlow and Keras to build a CNN-based classifier for identifying three common diseases: Black Rot, ESA (Net Blight), and Leaf Blight.
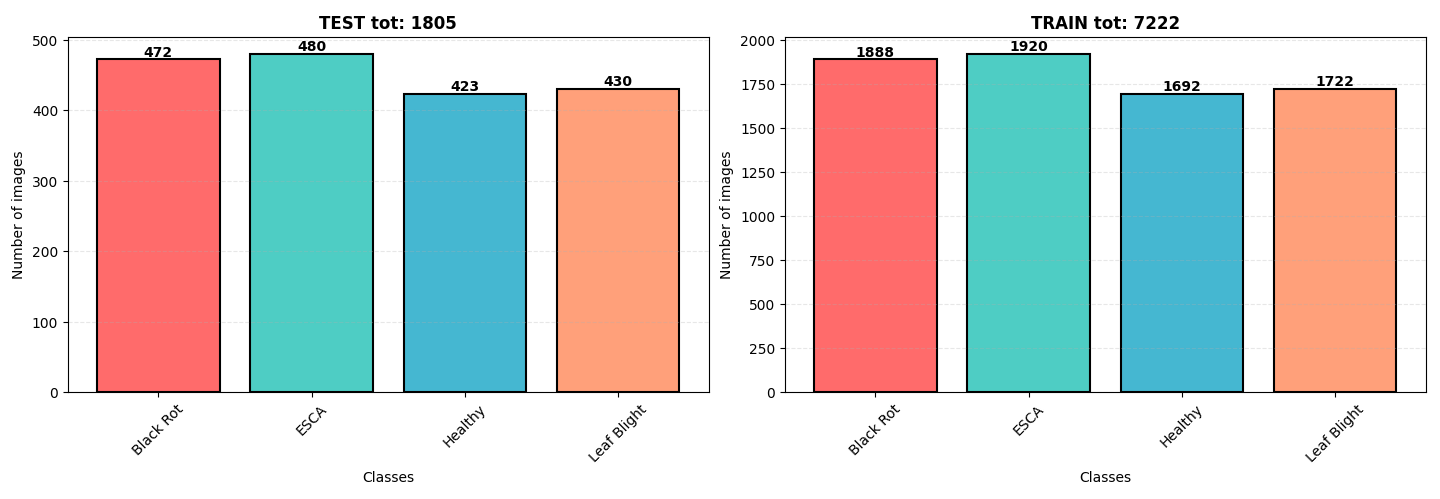
📁 Dataset
The dataset originates from Kaggle, containing 9,027 images of grapevine leaves. The diseases are categorized as:
- Black Rot
- ESCA (Net Blight)
- Leaf Blight
The dataset is well-balanced with a slight overrepresentation of ESCA and Black Rot. All images are in .jpeg format with dimensions 256x256 pixels.
📊 Model Architecture Selection
We evaluated pre-trained models from keras applications to balance accuracy, model size, and inference speed. The selection criteria included:
- Maximize accuracy
- Minimize size (1/size)
- Maximize CPU speed (1/CPU Time)
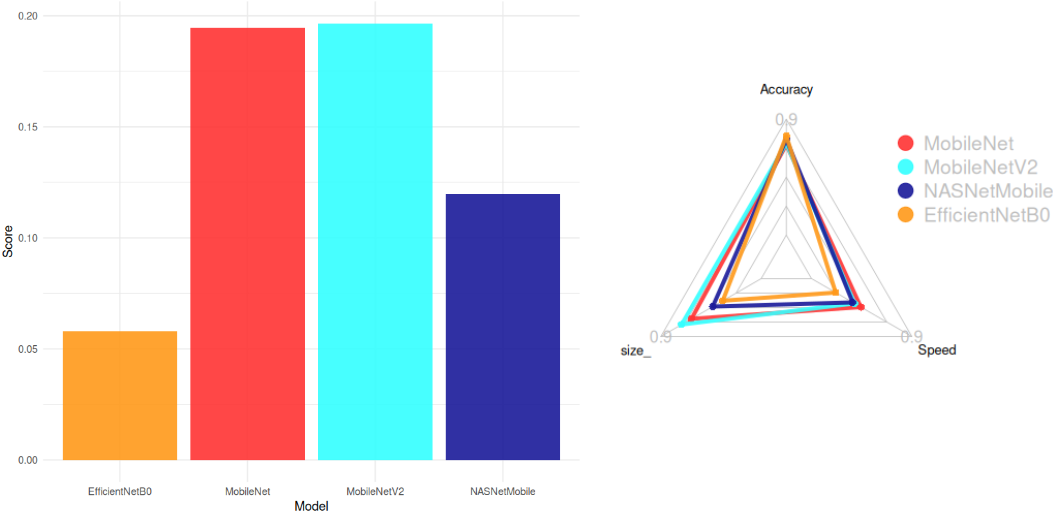
The score formula used for selection was:
Score = \frac{Accuracy}{Size . CPU Time}
Top Models (Score > 0.05):
- MobileNetV2 (Smallest: 14 MB, High Accuracy: 77%)
- MobileNet (Fastest: 22.6 ms)
- NASNetMobile
- EfficientNetB0
Conclusion:
MobileNetV2 was chosen for its optimal balance between accuracy, size, and speed.
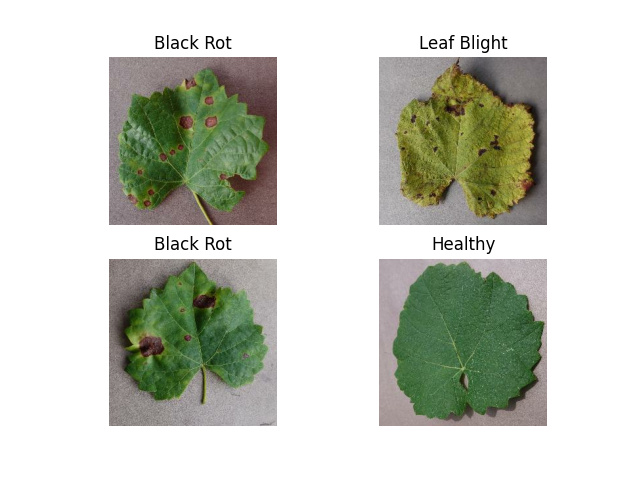
🍇 Grapevine Diseases
Key Diseases:
- Black Rot
- ESCA (Net Blight)
- Leaf Blight
🤖 Model Structure
Architecture:
# Auto Stop
early_stopping = EarlyStopping(monitor="val_loss", min_delta=0.2, patience=10)
# Model
model = Sequential()
model.add(tf.keras.applications.MobileNetV2(
input_shape=(IMG_HEIGHT, IMG_WIDTH, CHANNELS),
include_top=False,
weights='imagenet'
))
model.add(tf.keras.layers.GlobalAveragePooling2D())
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(NUM_CLASSES, activation='softmax'))
optimizer = tf.keras.optimizers.Adam(learning_rate=LEARNING_RATE)
model.compile(
optimizer=optimizer,
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy']
)
Parameters:
- Total params: 7.12M (27.17 MB)
- Trainable params: 2.36M (9.01 MB)
- Non-trainable params: 34.11K (133.25 KB)
🛠️ Training Details
- Batch Size: 32
- Epochs: 100 (reduced to 25 via early stopping)
- Data Augmentation: Not used (insufficient improvement in accuracy)
- Normalization: Pixel values normalized to [0, 1]
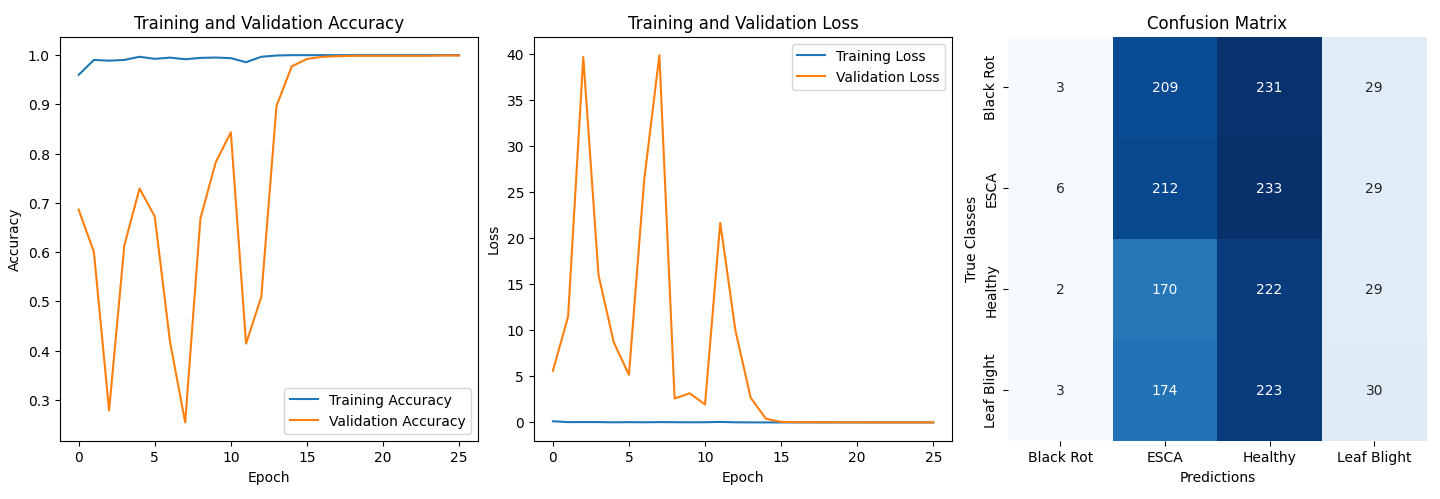
📊 Results
Performance:
- Validation Accuracy: ~99.9%
- Confusion Matrix Analysis:
- Model biased toward ESCA and Healthy classes.
- Suspected causes:
- Original dataset imbalance
- Similar visual features across diseases
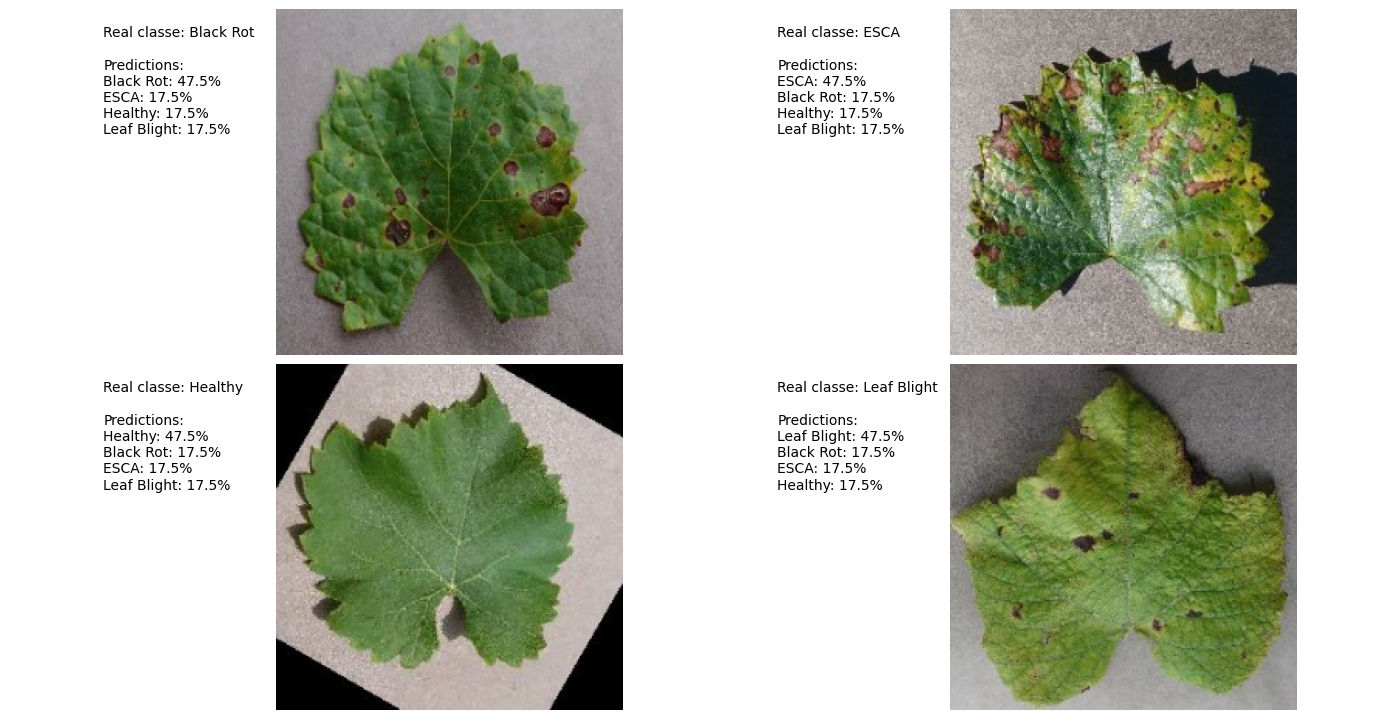
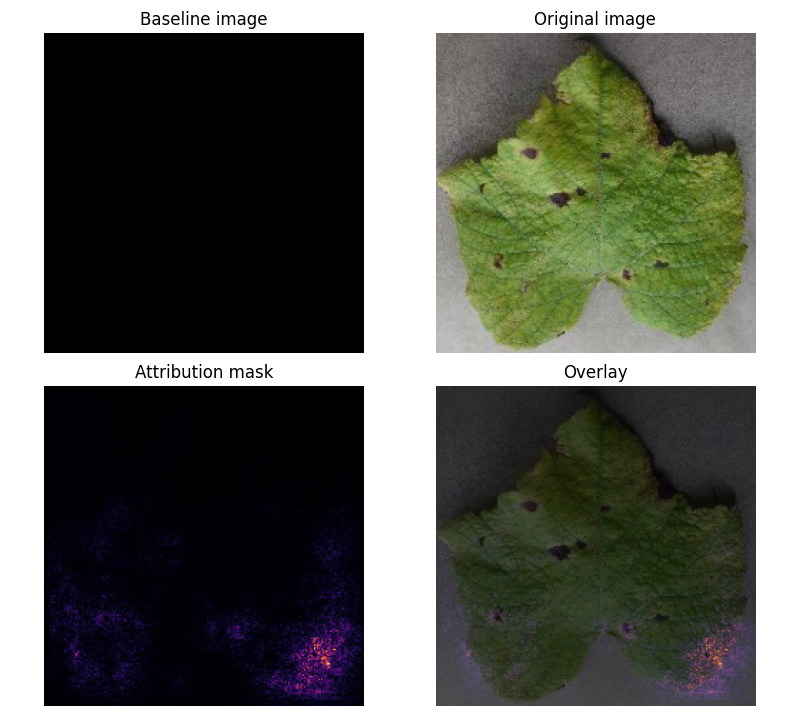
Prediction Example:
Attribution Mask:
- Key Insight: Model focuses on leaf shape rather than disease-specific features (e.g., black spots).
📚 ressources:
https://www.kaggle.com/code/ahmedmsaber/grape-leafs-diseases-mobilenetv2-val-acc-99
https://www.tensorflow.org/tutorials/images/classification?hl=en
https://www.tensorflow.org/lite/convert?hl=en
https://www.tensorflow.org/tutorials/interpretability/integrated_gradients?hl=en
🤖AI(s) : deepseek-coder:6.7b | deepseek-r1:8b