# Tensorflow Grapevine Disease Detection
## Description
This project develops a mobile application for detecting diseases on grapevines using a Deep Learning model. The implementation leverages TensorFlow and Keras to build a CNN-based classifier for identifying three common diseases:
Black Rot, ESA (Net Blight), and Leaf Blight.
## 📁 Dataset
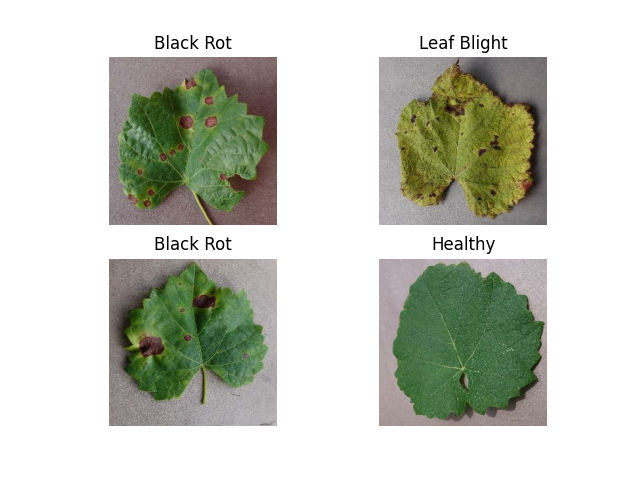
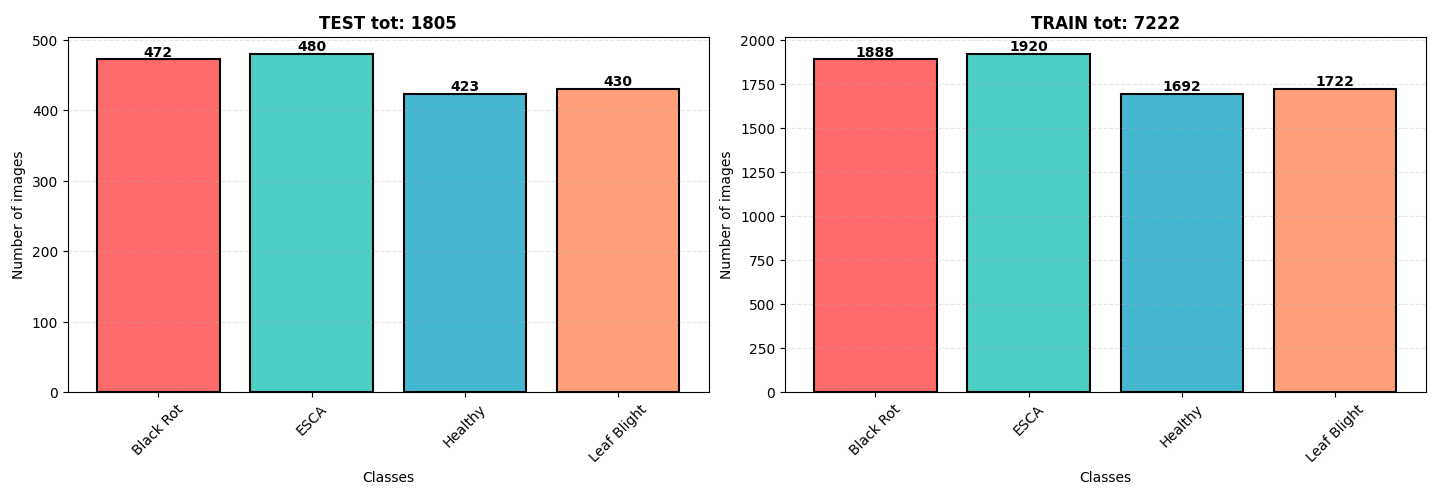
The dataset originates from [Kaggle](https://kaggle.com/datasets/rm1000/grape-disease-dataset-original), containing **9,027 images** of grapevine leaves. The diseases are categorized as:
- **Black Rot**
- **ESCA (Net Blight)**
- **Leaf Blight**
The dataset is **well-balanced** with a slight overrepresentation of **ESCA** and **Black Rot**. All images are in **.jpeg format** with dimensions **256x256 pixels**.

## 📊 Model Architecture Selection
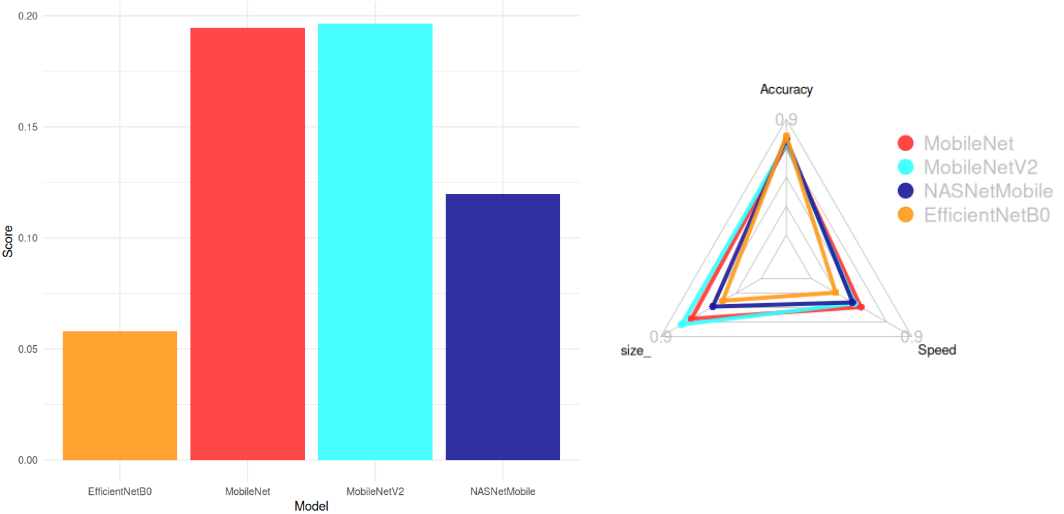
We evaluated pre-trained models from [`keras applications`](https://keras.io/api/applications/#usage-examples-for-image-classification-models) to balance **accuracy**, **model size**, and **inference speed**. The selection criteria included:
- **Maximize accuracy**
- **Minimize size** (1/size)
- **Maximize CPU speed** (1/CPU Time)
The **score formula** used for selection was:
$Score = \frac{Accuracy}{Size . CPU Time}$
**Top Models (Score > 0.05):**
1. **MobileNetV2** (Smallest: 14 MB, High Accuracy: 77%)
2. **MobileNet** (Fastest: 22.6 ms)
3. **NASNetMobile**
4. **EfficientNetB0**
**Conclusion:**
`MobileNetV2` was chosen for its optimal balance between accuracy, size, and speed.
## 🍇 Grapevine Diseases
### **Key Diseases:**
1. **Black Rot**
2. **ESCA (Net Blight)**
3. **Leaf Blight**
## 🤖 Model Structure
### **Architecture:**
```python
# Auto Stop
early_stopping = EarlyStopping(monitor="val_loss", min_delta=0.2, patience=10)
# Model
model = Sequential()
model.add(tf.keras.applications.MobileNetV2(
input_shape=(IMG_HEIGHT, IMG_WIDTH, CHANNELS),
include_top=False,
weights='imagenet'
))
model.add(tf.keras.layers.GlobalAveragePooling2D())
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(100, activation='relu'))
model.add(tf.keras.layers.Dense(NUM_CLASSES, activation='softmax'))
optimizer = tf.keras.optimizers.Adam(learning_rate=LEARNING_RATE)
model.compile(
optimizer=optimizer,
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy']
)
```
**Parameters:**
- **Total params:** 7.12M (27.17 MB)
- **Trainable params:** 2.36M (9.01 MB)
- **Non-trainable params:** 34.11K (133.25 KB)
## 🛠️ Training Details
- **Batch Size:** 32
- **Epochs:** 100 (reduced to 25 via early stopping)
- **Data Augmentation:** Not used (insufficient improvement in accuracy)
- **Normalization:** Pixel values normalized to [0, 1]
## 📊 Results
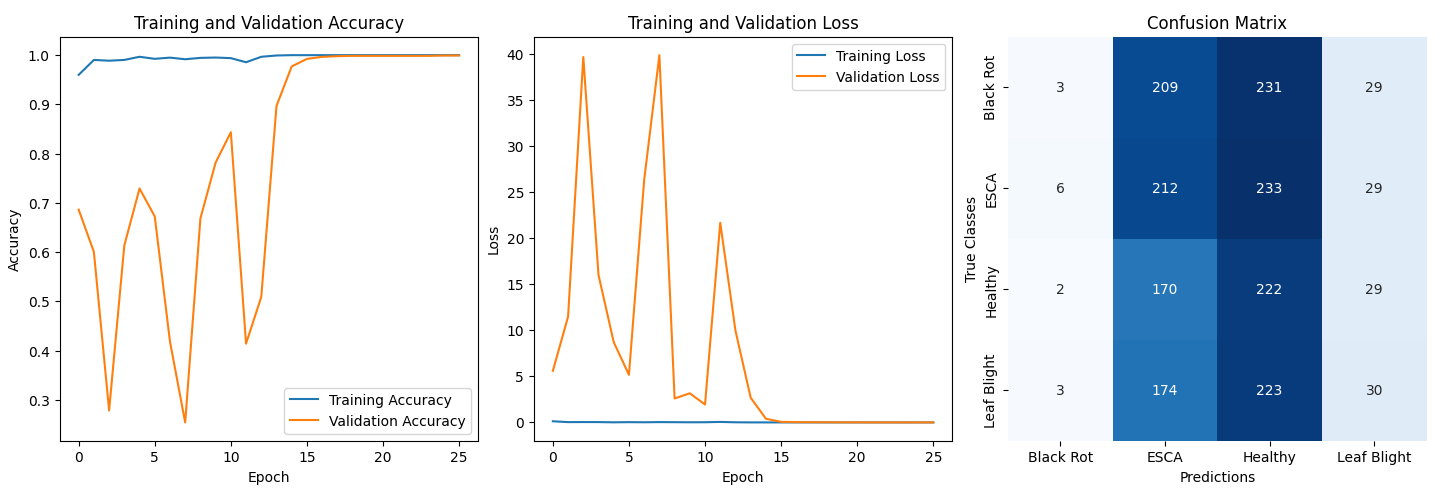
### **Performance:**
- **Validation Accuracy:** ~99.9%
- **Confusion Matrix Analysis:**
- Model biased toward **ESCA** and **Healthy** classes.
- Suspected causes:
1. Original dataset imbalance
2. Similar visual features across diseases
### **Prediction Example:**
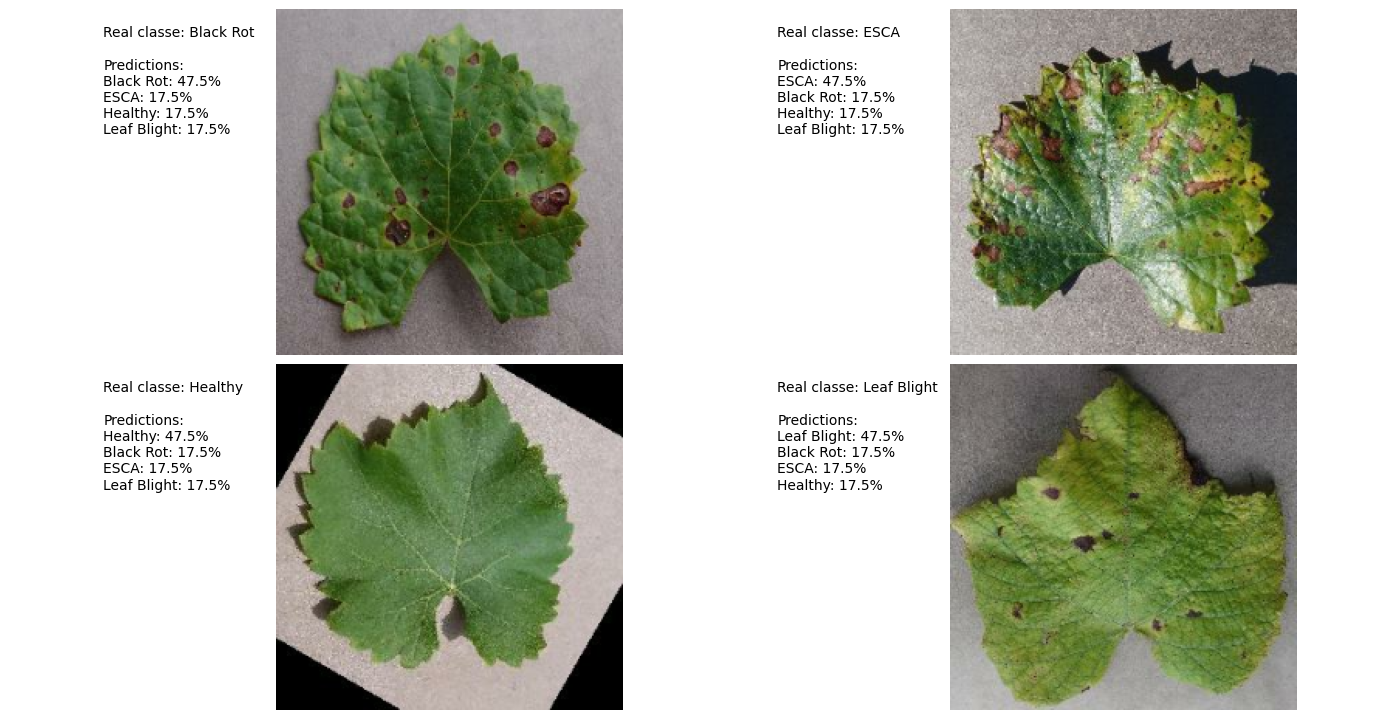

### **Attribution Mask:**
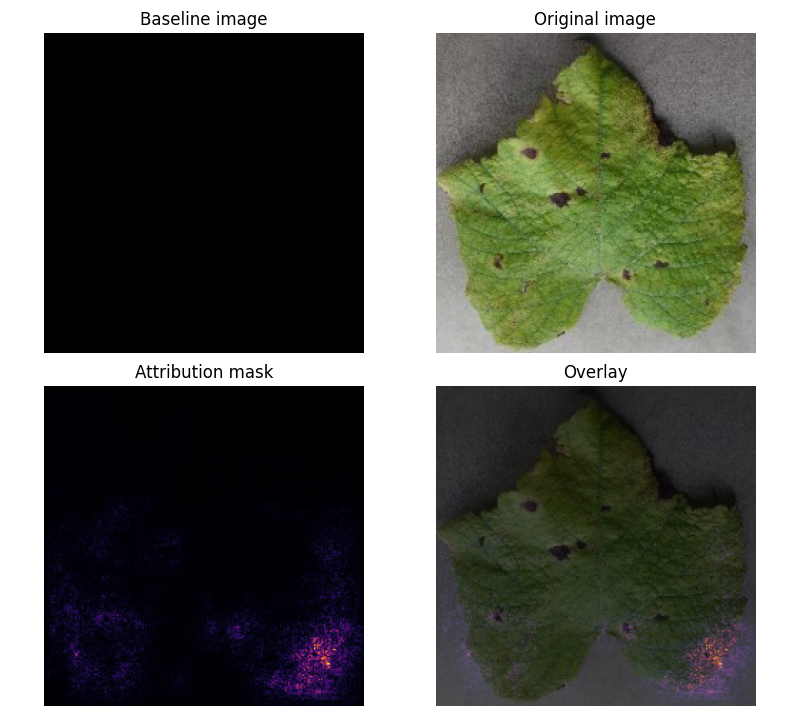
- **Key Insight:** Model focuses on leaf shape rather than disease-specific features (e.g., black spots).
### 📚 ressources:
https://www.kaggle.com/code/ahmedmsaber/grape-leafs-diseases-mobilenetv2-val-acc-99
https://www.tensorflow.org/tutorials/images/classification?hl=en
https://www.tensorflow.org/lite/convert?hl=en
https://www.tensorflow.org/tutorials/interpretability/integrated_gradients?hl=en
🤖AI(s) : deepseek-coder:6.7b | deepseek-r1:8b